Steve McDaniel, Brittney M. McInnis, Jonathan D. Hurt, , Lisa K. Kemp, Tyler W. Hodges12.11.19
Summary
Waterborne coatings, in particular, have been an important market for biocide manufacturers. However, the necessity to preserve these products from spoilage is challenging as regulations have increased on the type and level of biocide that can be used. This includes the use of formaldehyde releasers, growing restrictions on levels of isothiazolinones, as well as heavy metals such as chromium. Companies using these products have struggled to find ways to meet the new regulations while also offering quality, microbe-free products to their customers. The coatings industry’s focus on traditional biocides has led to predictions that no new actives are on the horizon, but bio-based additives offer solutions to turn that trend around. In cases where bioadditives alone are not sufficient to eliminate all microbial contamination, they can be used synergistically to lower the levels of traditional biocides needed. By applying biotechnology and molecular biology approaches, those levels can be lowered further. In addition, by capitalizing on the potential of these safe and effective bioadditives, manufacturers can expand into markets they might not have yet considered. Herein is presented the application of biotechnology for the preservation of coatings to achieve these goals.
Introduction
Waterborne coatings have been a lucrative market for biocide manufacturers over the last several decades with the move away from solvent-based systems and the increased demand for low-VOC coatings. With that move came an increased need to control microbial contamination and spoilage in water-based systems because these aqueous environments and organic nutrient sources offer an environmental niche well-suited for microbial growth. However, health concerns regarding the biocides used, coupled with increasing restrictions on the types and concentrations of traditional biocides, have increased the demand for development of novel and less toxic preservatives.1-3
Different microbial groups are responsible for negative effects on coating properties. For this reason, the biocides used routinely differ depending on their use in liquid in-can versus dry-film products. Bacteria are of greater concern for in-can preservation, while fungi and algae are the greatest threat to dry coatings, and both microbes and macrofauna are important when considering antifouling systems. In aqueous systems, microbial growth can impact the coating quality by hydrolyzing components, decreasing pH, producing gases, creating foul odors, discoloring the in-film product, and decreasing viscosity.4-5
Traditional biocides approved for in-can preservation typically include formaldehyde releasers, isothiazolinone derivatives, as well as brominated and other halogenated compounds. These are often combined with classes such as carbamates, quaternary amines, phenyl urea derivatives, and heavy metals for dry-film preservation and antifouling properties. Mechanisms of biocidal activity vary, but include effects due to alkylating agents, crosslinkers, electrophilic substances, membrane disruptors, as well as releasers of free radicals and reactive oxygen species.4 Some of these have more direct impact on humans (e.g., alkylating agents and crosslinkers), while others, such as the isothiazolinones, have lower direct toxicity to humans, but may cause sensitization following continued exposure.6-8 These and other health risks, as well as potential environmental impacts, have prompted many countries to restrict the levels of use and/or require special labeling.9-10
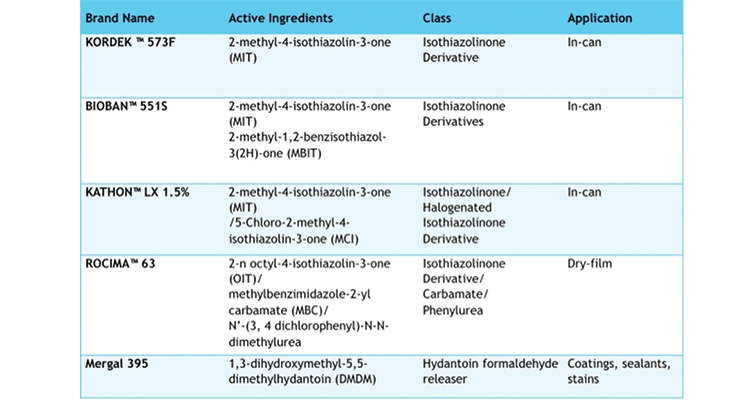
Table 1 lists some of the common traditional biocides. Many of the in-can preservative agents contain either 2-methyl-4-isothiazolin-3-one (MIT), or one of its isothiazolinone derivatives, making up brand name products like Kathon 1.5 and Rocima, all of which are subject to growing regulations restricting their usage and levels. From the health concerns with formaldehyde releasers and skin sensitizers, to the United States reaction following the increased standards in Europe [like REACH and the 2013 EU Biocides regulations (BPR) act], and calls by some to avoid biocide agents in building products altogether, these factors continue to put pressure on the industry.2, 11 Novel approaches are needed to meet these mounting regulatory hurdles, while still providing the needed protection in susceptible water-based coatings.
The use of rapid molecular techniques are described herein to select bio-based molecules which are safe and effective at controlling microbial contamination and can be used synergistically with existing biocides, with the goal of reducing the levels of such traditional biocides. Examples of bio-based molecules include enzymes like lysozyme and glucose oxidase, small peptides similar in activity to natural antimicrobial peptides like defensins, but smaller in molecular weight, and even whole cells containing such molecules. By retaining the natural biological function (in either direct antimicrobial activity or synergistic activity with a traditional biocide) it is possible to utilize these molecules in liquid and dry film coating systems.
Experiments and Results
Application of Molecular Techniques to Screen Bio-based Additives as In-Can Preservatives
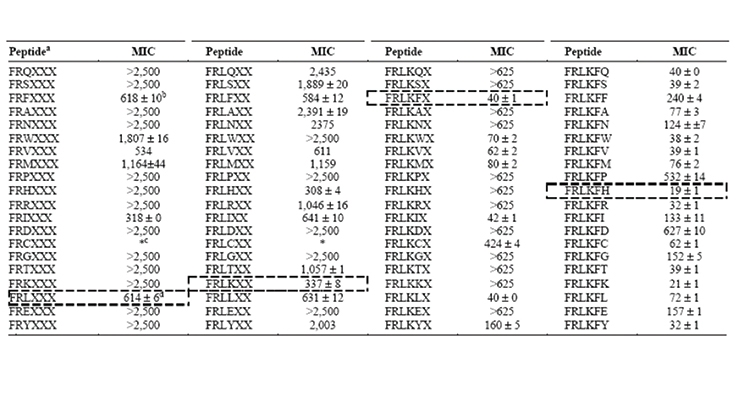
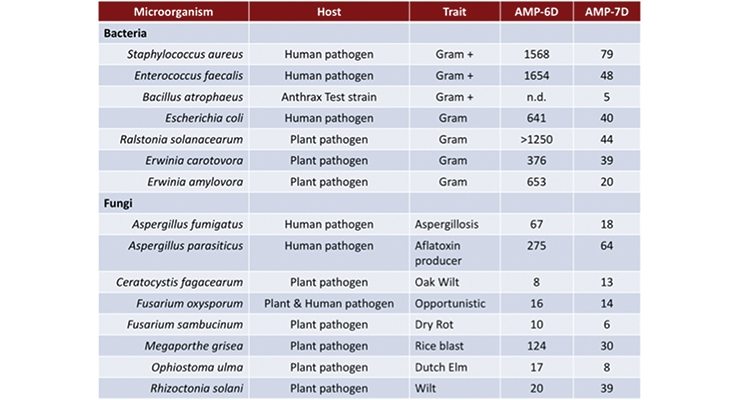
The small peptides, AMP-6 and AMP-7, are 6-7 amino acids in length (molecular weight under 1000 g/mol) and were selected from a peptide library due to their antimicrobial properties. The library was created using combinatorial synthesis methods to produce over 53 million possible peptides that could be screened for specific antimicrobial activities. Figure 1A shows an example of the iterative amino acid substitution process used to generate the library and minimum inhibitor concentrations (MIC) against Fusarium, a common filamentous fungus.12-16 By altering the peptide sequence, it is possible to “tune” the activity towards specific target microbe groups like bacteria versus fungi and molds. Using the combinatorial chemical approach, progressively lower concentrations are needed to kill the same target microorganism as the peptide chemical structure becomes more defined (which in this case is a 6-amino acid chain). In the case of AMP-6 (a 6-amino acid peptide), greater activity was seen against fungi (as indicated by a lower MIC number) than the bacteria tested (a higher concentration needed). With the addition of just a single amino acid, AMP-7 shows satisfactory activity against both bacteria and fungi (Figure 1B).
Molecular Biology Techniques Drop Test Times to Minutes vs. Weeks
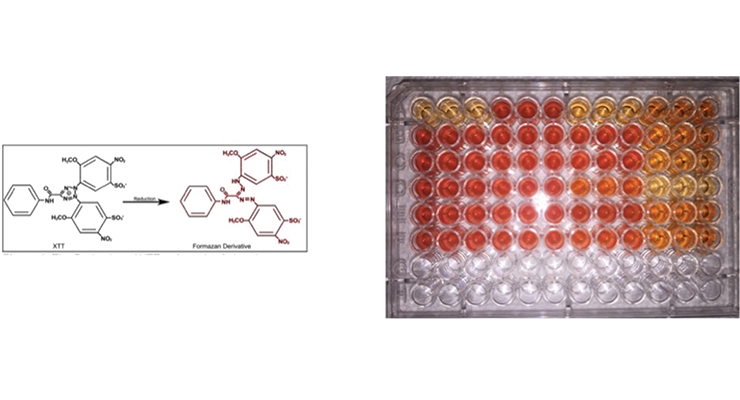
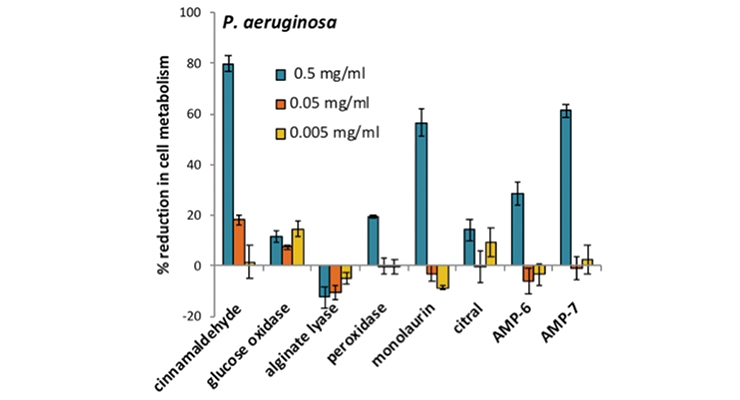
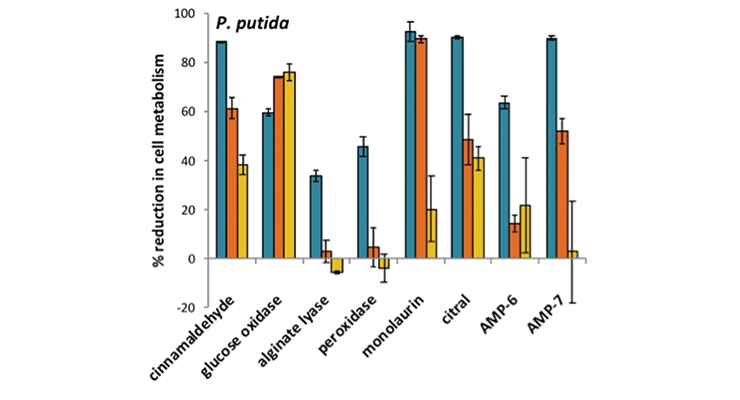
The use of molecular-based microbial viability assays, such as the XTT assay, allows for rapid assessment of the effects of candidate antimicrobials.17 XTT is a tetrazolium dye (2,3-bis-(2-methoxy-4-nitro-5-sulfophenyl)-2H-tetrazolium-5-carboxanilide) that is reduced to colored products by metabolically active cells (darker reddish-brown wells) versus cells that are heat-killed or treated with ethanol, which remain the lighter yellow color (Figure 2A). The difference in the absorbance spectrum can be measured using a spectrophotometer and the viability of the cell population rapidly assessed. Applying these assays, it is possible to screen for an antimicrobial’s selectivity against particular contaminating microorganisms rather than apply broadly toxic biocides. Figure 2B shows that the concentrations of bioadditives needed to control different contaminating strains of Pseudomonas can vary drastically. For P. putida, several agents were effective at various concentrations (as indicated by higher % reduction in metabolism), while P. aeruginosa required the higher test concentration (blue bars) of the tested bio-additive.
Synergistic Activity with Existing Traditional Biocides
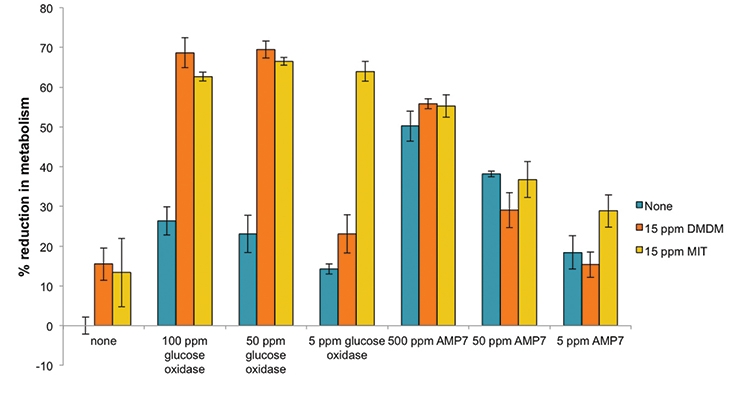
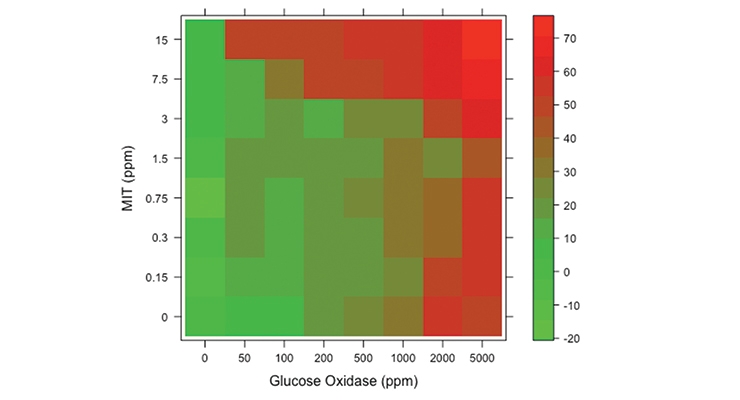
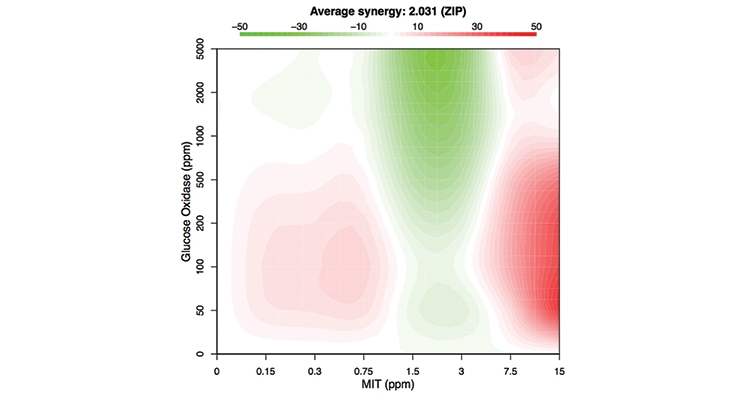
Based on the initial XTT screening results, two common biocides were evaluated for synergistic activity with these bio-based agents: an isothiazolinone (MIT), and the formaldehyde releaser, 1,3-dihydroxymethyl-5,5-dimethylhydantoin (DMDM) (Figure 3). A mixed inoculation test panel of six common bacterial spoilage agents, including several Pseudomonads and other gram-negatives like Enterobacter and a gram-positive Bacillus, was used to evaluate the bioadditives and biocides. In the first column, these agents are shown to be effective alone at 15 ppm (15 ppm is the cutoff for labeling as a skin sensitizing agent versus the more stringent EUH208 “May produce an allergic reaction”). At 15 ppm little inhibition was seen in this assay (only around 15% reduction in metabolism), but when combined with particular bio-based additives, an increase in the effectiveness was seen (from 15% up to over 70%, which is comparable to what is seen in heat-killed cells). Particularly with the glucose oxidase synergistic activity an effective kill rate at 15 ppm is achieved. Glucose oxidase catalyzes the oxidation of glucose and subsequent release of hydrogen peroxide that can induce cellular damage.18, 20 Glucose oxidase is listed by the Food and Drug Administration as generally recognized as safe (GRAS) for various intended uses and is frequently used in the food industry.19 It is produced on commercial scale and available in bulk quantities. This apparent synergistic activity was confirmed by testing varying concentrations of glucose oxidase and MIT in the XTT assay. The responses of the test panel bacteria to various combinations of glucose oxidase and MIT are displayed in the heatmap (Figure 4A), which shows the relative response as a color from red (highest percent growth inhibition) to green (lowest percent growth inhibition). To test whether antagonism or synergy exists between these two compounds, the responses from glucose oxidase and MIT combinations were used to determine the synergy score using the zero interaction potency (ZIP) method, which returns a score based on the deviation of the actual response from the expected response.21 These scores are visualized for each combination in a contour plot (Figure 4B), which suggests that the two antimicrobials generally interact in a synergistic manner. Regions of synergism exist for concentrations of glucose oxidase between 50 ppm and 500 ppm and for concentrations of MIT ranging from below 0.15 ppm to 1 ppm, as well as the greatest synergy seen for concentrations of glucose oxidase between 10 ppm and 1,000 ppm and for concentrations of MIT ranging from below 7 ppm to 15 ppm. This suggests that the glucose oxidase and MIT are complementing each other’s function at both low and high concentrations allowing for lower levels of MIT to show effective antimicrobial activity.
Evaluation of Glucose Oxidase Synergy with Traditional Biocides Using ASTM D2574
The antimicrobial activity of glucose oxidase, detected using the rapid molecular techniques, was further tested as an in-can coatings preservative using the traditional testing method, ASTM D2574 Paint Spoilage standard.22 By using the above molecular methods to rapidly screen target microbes, it is possible to select the most promising candidates to classic tests like ASTM D2574.
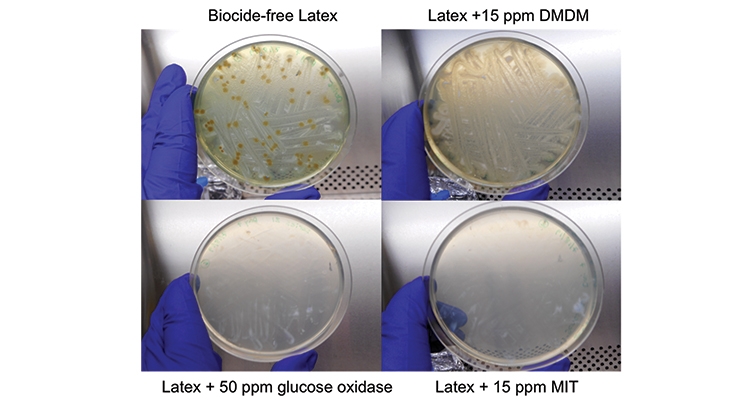
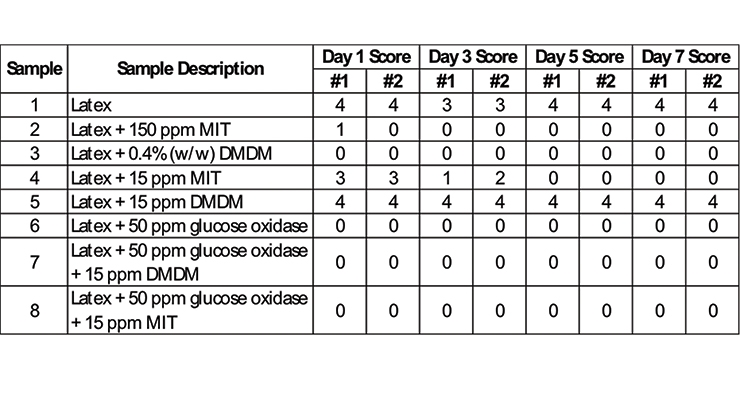
For this testing, a biocide-free acrylic latex was used (see Appendix - Materials and Methods for formulation). The same spoilage agent test panel used for the XTT assay was used to challenge the acrylic latex; the samples were monitored over seven days. The selected dosage of glucose oxidase for the ASTM D2574 procedure was based on the results from the XTT assay, where 50 ppm showed synergy with both MIT and DMDM while 5 ppm was only synergistic with DMDM. Figure 5 shows example plates from bio-based additive versus traditional biocides on the final day. Over the 7-day monitoring period the biocide-free latex controls scored a 4 for each day (complete overgrowth).
While the recommended dosages of MIT and DMDM achieved a 0 (no growth) on day 1, the lower doses of 15 ppm were less effective with DMDM still showing a 4 on day 7 and MIT not achieving a 0 until day 5. However, in the ASTM method using the biocide-free latex, the lowest test concentration of glucose oxidase (50 ppm) achieved a 0 on day 1 and when combined with 15 ppm MIT or DMDM a 0 was also achieved on day 1 (Table 2).
While it is a positive result that glucose oxidase alone showed such high activity, further studies are being conducted at lower concentrations of glucose oxidase in order to characterize any synergistic activity with traditional biocides in the acrylic latex.
Molecular Approaches for Detecting and Eliminating Paint Spoilage Agents
While classic testing methods like ASTM D2574 identify broadly toxic agents that kill all challenge organisms, they fail to selectively identify agents effective against particular strains. By using DNA sequencing methods, it is possible to identify specifically which bacteria are killed by particular bio-based additives for precision treatments (Figure 6A).
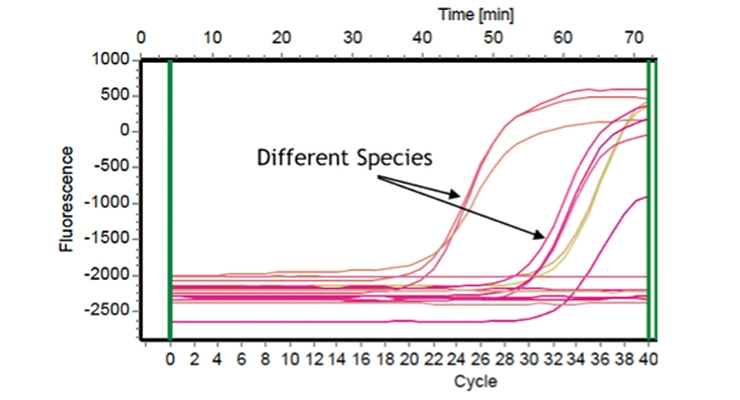
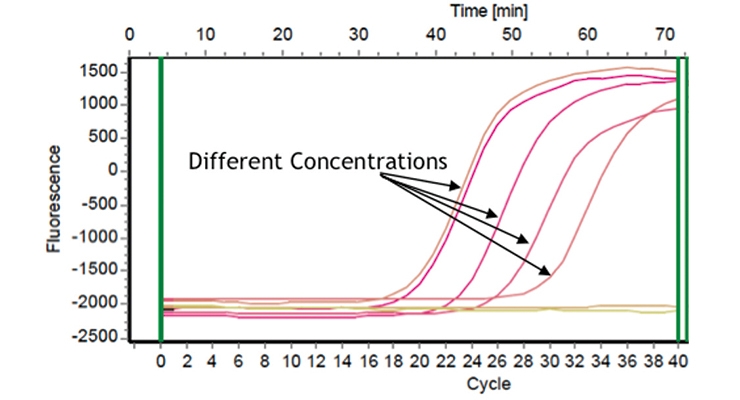
Real-time quantitative polymerase chain reaction (PCR) was used for direct analysis of paint for microbial contamination and identification of the specific contaminant. The graph in Figure 6B shows the use of selective probes to identify particular bacterial species. The fluorescence increases as the strain-specific probes successfully amplify the target DNA. These methods can also be used to determine the levels of contamination as well. Figure 6C shows a dose response curve using Enterobacter, which indicates the changes in cycle number needed to see amplification at the different concentrations. By identifying the particular level of bacterial contamination and treating accordingly, the need to use high levels of broadly toxic biocides in products decreases.
Expanded Markets Using Bio-based Additives: Food Safety and Combating Antibiotic Resistance
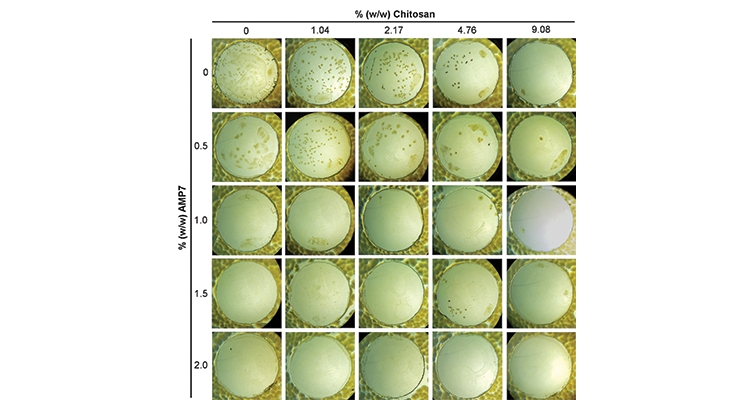
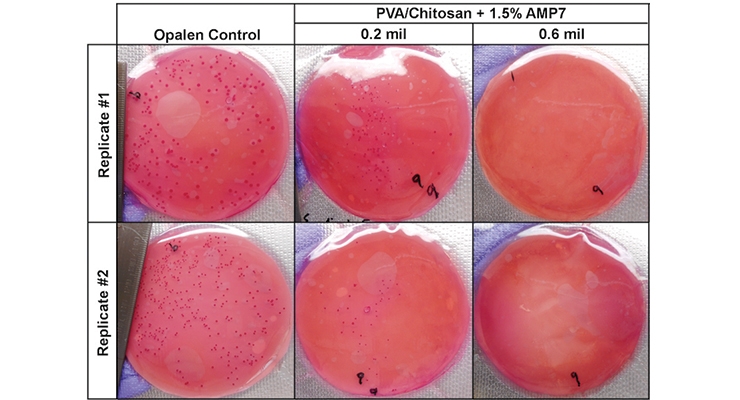
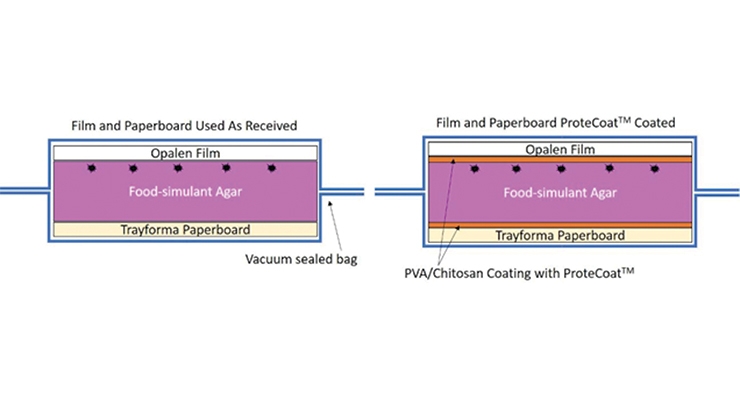
With the low toxicity bio-based additives, some of which have GRAS status for various applications, manufacturers can expand into new markets beyond in-can coating preservation - one example being food packaging. A rapid screening technique based on inhibition of microbial colony formation on agar plates was used to find active agents against E. coli and explore any potential synergy between active agents.23 Figure 7A shows an image of small 2.5 cm polyethylene terephthalate (PET) cutouts coated with polyvinyl alcohol (PVA) incorporating bioadditives used to test for contaminant inhibition (in this case E. coli) on agar plates. With increasing concentrations of each agent alone, inhibition of growth was observed, as measured by the reduction in the number of visible colonies under the plastic. By combining the two agents, a further reduction in the concentration of each agent needed was achieved. Following these results, a lab-scale test was developed using food simulant patties (microbial growth media with cell growth color indicator) that can be challenged with the test microbe and then vacuum sealed with either bio-additive coated sealing plastic itself or sandwiched between inserts with bio-additive coatings to prevent foodborne contamination (Figure 7C). As seen in Figure 7B, at an optimal dose of bio-additive in the packaging coating, the viability of the E. coli is reduced to zero (no colonies).
Other examples of expanded markets include products to help combat a growing global healthcare concern: antibiotic resistance. Not only is there a need for safe and effective antimicrobials (such as the bio-based agents described above), but also methods to combat transfer of antibiotic resistance (as a cell is lysed, the DNA spills out and can be taken up by other cells).
Enzymatic coating surfaces have been successfully created that can breakdown and destroy cell- free DNA spilled from a cell before it can be taken up by other cells.24 The top left panel in Figure 8 shows the effective killing of E. coli by AMP-7 (3% total solids), as indicated by lack of bacterial colony growth around the edges of the AMP-7 polyurethane cutout versus the control when placed on contaminated agar plates. The middle panel shows the destruction of DNA by a polyurethane coating containing the nuclease, DNase I (1.5% total solids). The DNA used here carries a gene for resistance to ampicillin, a beta-lactam antibiotic related to penicillin. The DNA was applied to the surface and allowed to dry. Following exposure, the DNA was resuspended from the coating surface and analyzed. On both ends of the image in Figure 8 are three wells each of coating controls (either the substrate itself or the polyurethane coating). In the middle are wells with the DNA exposed to the nuclease-containing coating. Distinct bands are no longer detected when exposed to the nuclease-containing coating surface, though the smear of the smaller breakdown products can be seen, indicating that the DNA has been destroyed. Furthermore, on the right in Figure 8, it was demonstrated that following exposure of the ampicillin resistance gene containing DNA to the nuclease-coated surface, no transfer of antibiotic resistance to E. coli occurs. The agar plates in this image have ampicillin added to them, and only E. coli that were able to take the DNA directly from the surface are able to grow (n=3: 36, 37, and 26 colonies). However, E. coli from the nuclease-coated surface did not develop resistance to ampicillin because the DNA was destroyed, resulting in no colony growth (n=3: 0, 0, and 0 colonies). The bottom of Figure 8 sums up the overall process. After the bacteria are lysed and killed by AMP-7, the free DNA is degraded by nucleases in the coating, and the resistance gene is not passed.
Further Coating Manufacturing Applications to Reduce Biocide Levels
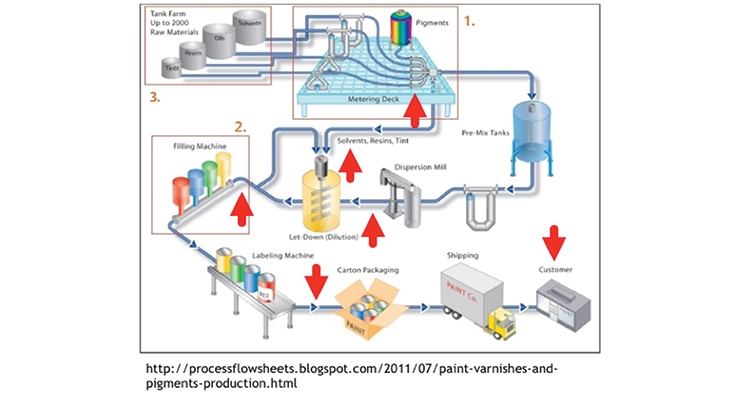
In the coatings manufacturing arena these technologies can be applied to expand applications and market areas even further, such as surgical strikes by inclusion of selected bioadditives along the coating manufacturing process line to decrease the need to incorporate high levels of biocides in the final product (Figure 9A). By using the previously mentioned molecular methods, the source contaminant can be rapidly identified and equipment lines and materials treated accordingly with bio-additives at the point of contamination. By utilizing biomolecules, like enzymes, and the tunability of AMPs, the targeted use of these agents within the plant becomes possible. That protection can potentially continue from the plant to the consumer without an added in-can biocide by coating the can itself with bio-additive protectants as shown in Figure 9B. Greater access to oxygen at the walls and the paint/headspace interface makes these active sites of microbial growth, and condensation buildup on the lid makes this another potential problem site to target.
Conclusion
In conclusion, given the growing regulatory challenges facing traditional biocide usage, alternatives are needed to 1) address the increased regulations and 2) expand upon existing markets. One such approach to address those issues is the use of rapid molecular methods to identify bio-based additives such as peptides and enzymes for selective contaminant targeting. By using these methods, agents were identified that have either inherent antimicrobial activity, like AMP-6 and AMP-7, or GRAS enzymes like glucose oxidase. In instances where the bio-based biocide is not broadly antimicrobial on its own, it is shown here that such bio-based molecules facilitate the activity of traditional biocides like MIT and allow for lower levels of biocide use.
Acknowledgements
Aayushma Kunwar, Kyle Wilhelm and Earnest Couch, student interns from William Carey University, Hattiesburg, MS, are thanked for their assistance with these projects.
References
1. Cresswell, T.; Richards, J. P.; Glegg, G. A.; Readman, J. W., The impact of legislation on the usage and environmental concentrations of Irgarol 1051 in UK coastal waters. Marine Pollution Bulletin 2006, 52 (10), 1169-1175.
2. Brinch, A.; Hansen, S.; Hartmann, N.; Baun, A., EU regulation of nanobiocides: challenges in implementing the biocidal product regulation (BPR). Nanomaterials 2016, 6 (2), 33.
3. White Paper: Antimicrobial Building Products Should Be Avoided Whenever Possible. http://perkinswill.com/news/perkinswill-white-paper-antimicrobial-building-products-should-be-avoided-whenever-possible (accessed December 2017).
4. Sauer, F., Microbicides in Coatings. Vincentz Network: Hanover, Germany, 2017.
5. Winkowski, K., Efficacy of in can preservatives. European coatings journal 2001, (1-2), 87-91.
6. Chan, P. K.; Baldwin, R. C.; Parsons, R. D.; Moss, J. N.; Stiratelli, R.; Smith, J. M.; Hayes, A. W., Kathon biocide: manifestation of delayed contact dermatitis in guinea pigs is dependent on the concentration for induction and challenge. Journal of investigative dermatology 1983, 81 (5), 409-411.
7. Schwensen, J. F.; Menné, T.; Andersen, K. E.; Sommerlund, M.; Johansen, J. D., Occupations at risk of developing contact allergy to isothiazolinones in D anish contact dermatitis patients: results from a D anish multicentre study (2009–2012). Contact Dermatitis 2014, 71 (5), 295-302.
8. Schwensen, J. F.; Lundov, M. D.; Bossi, R.; Banerjee, P.; Gimenez-Arnau, E.; Lepoittevin, J. P.; Lidén, C.; Uter, W.; Yazar, K.; White, I. R., Methylisothiazolinone and benzisothiazolinone are widely used in paint: a multicentre study of paints from five E uropean countries. Contact Dermatitis 2015, 72 (3), 127-138.
9. Bollmann, U. E.; Fernández-Calviño, D.; Brandt, K. K.; Storgaard, M. S.; Sanderson, H.; Bester, K., Biocide runoff from building facades: degradation kinetics in soil. Environmental science & technology 2017, 51 (7), 3694-3702.
10. Lundov, M. D.; Kolarik, B.; Bossi, R.; Gunnarsen, L.; Johansen, J. D., Emission of isothiazolinones from water-based paints. Environmental science & technology 2014, 48 (12), 6989-6994.
11. REACH Legislation. https://echa.europa.eu/regulations/reach/legislation (accessed 08/09/19).
12. McDaniel, C.S., Anti-fouling Coatings and Coatings,
U.S. patent application 14/097,128, July 9, 2015.
13. McDaniel, C.S., Antifungal Coatings and Coatings, U.S. patent 8,497,248, July 30, 2013.
14. McDaniel, C.S., Coating Compositions having Peptidic Antimicrobial Additives and Additives of Other Configurations, U.S. patent 8,618,066, December 31, 2013.
15. McDaniel, C.S., Antifungal Coatings and Coatings, U.S. patent 7,932,230, April 26, 2011.
16. McDaniel, C.S., Anti-fouling Coatings and Coatings, U.S. patent 7,939,500, May 10, 2011.
17. XTT Cell Proliferation Assay Kit Instruction Manual. American Type Culture Collection: Manassas, VA, 2011.
18. Hodges, T. W.; Kemp, L. K.; Mcinnis, B. M.; Hurt, J. D.; McDaniel, S.; Rawlins, J. W.; Wilhelm, K. L., Proteins and Peptides: as Replacements for Traditional Organic Preservatives: Part I. CoatingsTech 2018, 15 (4).
19. Bankar, S. B.; Bule, M. V.; Singhal, R. S.; Ananthanarayan, L., Glucose oxidase—an overview. Biotechnology advances 2009, 27 (4), 489-501.
20. Tiina, M.; Sandholm, M., Antibacterial effect of the glucose oxidase-glucose system on food-poisoning organisms. International Journal of Food Microbiology 1989, 8 (2), 165-174.
21. He, L.; Kulesskiy, E.; Saarela, J.; Turunen, L.; Wennerberg, K.; Aittokallio, T.; Tang, J., Methods for High-throughput Drug Combination Screening and Synergy Scoring. Methods Mol Biol 2018, 1711, 351-398.
22. ASTM Standard D2574-16, Standard Test Method for Resistance of Emulsion Coatings in the Container to Attack by Microorganisms. ASTM International: West Conshohocken, PA, 2016.
23. McInnis, B. M.; Hodges, T. W.; Kemp, L. K.; Hurt, J. D.; McDaniel, S.; Kunwar, A., Bio-based Antimicrobial Food Packaging Coatings. CoatingsTech 2018, 15 (9), 36-43.
24. Ellison, C. K.; Dalia, T. N.; Vidal Ceballos, A.; Wang, J. C.-Y.; Biais, N.; Brun, Y. V.; Dalia, A. B., Retraction of DNA-bound type IV competence pili initiates DNA uptake during natural transformation in Vibrio cholerae. Nature Microbiology 2018, 3 (7), 773-780.
25. Reed, J. D.; Edwards, D. L.; Gonzalez, C. F., Synthetic Peptide Combinatorial Libraries: A Method for the Identification of Bioactive Peptides Against Phytopathogenic Fungi. Molecular Plant-Microbe Interactions 1997, 10 (5), 537-549.
26. ZymoBIOMICS DNA/RNA Miniprep Kit. www.zymoresearch.com: Ver. 1.2.0.
27. Lee, C. S.; Wetzel, K.; Buckley, T.; Wozniak, D.; Lee, J., Rapid and sensitive detection of Pseudomonas aeruginosa in chlorinated water and aerosols targeting gyrB gene using real-time PCR. Journal of applied microbiology 2011, 111 (4), 893-903.
28. Preston, K. E.; Radomski, C. C.; Venezia, R. A., Nucleotide Sequence of the ChromosomalampC Gene of Enterobacter aerogenes. Antimicrobial agents and chemotherapy 2000, 44 (11), 3158-3162.
Appendix Materials and Methods
Antimicrobial Peptide Library
See materials and methods in Reed et al. 200725 and patent numbers patent numbers 5,602,097, 5,885,782, and 6,020,312.
XTT Assay
See materials and methods in Hodges et al. 2018.18
XTT Assay for Synergy with MIT and DMDM
See materials and methods in Hodges et al. 2018.18 For experiments involving the combination glucose oxidase and MIT, the interaction between the two additives (i.e., synergistic, additive, antagonistic) was evaluated using the zero interaction potency (ZIP) model with the R package synergyfinder.21,23
ASTM D2547 Using Acrylic Latex
Antimicrobial activity of traditional and bio-based biocides were assessed using the ASTM method D254722 using a custom acrylic latex (50 wt% solids, pH = 8, Tg = 18 °C) without biocide. Twenty-five milliliters of liquid acrylic latex was dispensed into sterile glass jars, and appropriate quantities of glucose oxidase, MIT, or DMDM were hand-stirred into the coating using sterile glass rods. Sterile swabs were dipped into each sample and used to streak Tryptic Soy Agar (TSA) plates to ensure that the samples were free of bacterial contamination. Bacterial cultures of Pseudomonas aeruginosa, Pseudomonas putida, Pseudomonas fluorescens, Bacillus cereus, Alcaligenes faecalis, and Enterbacter aerogenes were grown in 5 mL Tryptic Soy Broth (TSB) at 30 °C with agitation for 24 hours. Sterile loops full of each culture were passed into new 5 mL of TSB, which were incubated at 30 °C with agitation for 24 hours. Equal volumes of the cultures were passed once more, with a final incubation at 30 °C with agitation for 24 hours. The cultures were mixed, and 250 µL of the bacterial mixture was dispensed into each latex sample. Sterile swabs were dipped into each sample and used to streak duplicate TSA plates 1, 3, 5, and 7 days following inoculation. The TSA plates were incubated at 30 °C for 7 days, after which each plate was scored using the score system mandated by the ASTM
D2547 procedure:
0 - no bacterial recovery
1 - trace contamination (1 to 9 colonies)
2 - light contamination (10 to 99 colonies)
3 - moderate contamination (> 100 distinct colonies)
4 - heavy contamination (continuous smear of growth,
colonies have grown together and are indistinguishable)
Isolate Identification using Molecular Analysis
Following inoculation of biocide-free acrylic latex paint or bio-based additive treated paint according to ASTM D2547, bacterial colonies were steaked for isolation on TSB agar. Isolated colonies were transferred into microcentrifuge tubes containing 100 µL of chelex solution (BioRad Instagene Matrix). The tubes were vortexed for 15 seconds and placed in a 70 °C water bath for 1 hour, with vortexing repeated every 15 minutes. Samples were then centrifuged at 12,000xg for 2 min and either used directly in polymerase chain reaction (PCR) or frozen. PCR primers targeting a portion of the 16S ribosomal RNA (rRNA) gene were used, specifically 8F (5’-AGAGTTTGATCMTGGCTCAG-3’) and 1392R (5’-ACGGGCGGTGTGTACA-3’). GE Healthcare Illustra PureTaq PCR beads were used for PCR amplification with the following conditions: Per reaction vial: 2 µL 8F primer, 2 µL 1392R primer, 2 µL template DNA, and 19 µL sterile water. Thermal cycler conditions were 94 °C for 5 min followed by 30 cycles of 94 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min followed by 72 °C for 7 min. Following completion of PCR, Invitrogen ChargeSwitch-Pro PCR Cleanup Kit was used to purify the amplification products. PCR products containing the amplified region of the 16S rRNA gene were then sent for sequencing at Eurofins Genomics (Louisville, KY). Upon receipt of the sequence data, sequences were searched against the NCBI Genbank using BLAST (Basic Local Alignment Search Tool). This allows for strain identification based on sequence similarity to deposited 16S rRNA gene sequences.
Molecular Paint Analysis Using Quantitative Real-time PCR
Direct DNA extraction from paint: Five types of commercially available DNA extraction kits were tested for effectiveness in isolating amplifiable bacterial DNA from latex acrylic paint (AxyPrep Mag Tissue-Blood gDNA, Invitrogen PureLink, Microbiome DNA Purification Kit, Invitrogen PureLink Genomic DNA Mini Kit, and ZymoBiomics DNA Mini Kit). Off-the-shelf Sherwin Williams Harmony EgShel was inoculated with overnight cultures of set bacteria, in this example Enterobacter aerogenes, followed by DNA extraction using the commercially available kits.26 Amplification using universal bacterial 16S primers (16F and 16R) and standard end-point PCR showed amplification as indicated by appropriate size bands in gel electrophoresis.
Reaction Conditions and PCR Primers: Quantitative Real-time PCR conditions were optimized using primers targeting functional genes in three species from the ASTM challenge method (Pseudomonas aeruginosa, Enterobacter aerogenes, and Bacillus cereus). Where necessary, primers were designed using the National Center for Biotechnology Information Primer-BLAST function. An Eppendorf RealPlex thermal cycler and SYBR Green were used for detection of amplified DNA (Applied Biosystems™ PowerUp™ SYBR™ Green Master Mix). Primers were synthesized by Eurofin Operon (Louisville, KY).
Primers sequences used:
Pseudomonas aeruginosa: gyrB gene (DNA gyrase, subunit B)27
PA-722 primer set: 722F:5’-GGCGTGGGTGTGGAAGTC-3’
and 722R: 5’-TGGTGGCGATCTTGAACTTCTT-3’)
Enterobacter aerogenes: Designed based on beta-lactamase
AmpC gene28
EA-PP5 primer set: PP5F:5’-GACTACGCCTGGGGTTATCG-3’
PP5R: 5’-CGCTGTTCATATTCGCCAGC-3’
Typical reaction mixtures: Primers were diluted to 20 µM stocks in sterile ddH20 and 0.5 µL added per 20 µL reaction for a final concentration of 500 nM. DNA concentrations varied between experiments: extracted genomic DNA was used at 2 µL in initial experiments and, if quantitated, it was added at 0.5 µL per 20 µL reaction (pg or ng range). Example of a typical 20 µL reaction setup is as follows:
Per 20 µL reaction
0.5 µL Forward Primer
0.5 µL Revers Primer
0.5 µL DNA (up to 2 µL)
7.5 µL sterile H2O
10 µL SYBR Green Master Mix
Thermal Cycler Conditions: 50 °C for 2 min, 95 °C for 2 min followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min with fluorescence measured at 520 nM at the end of each cycle. Cycle threshold values (ct values) were calculated automatically by the instrument. This is the number of cycles taken for the fluorescence signal to cross background noise level and is inversely proportional to the amount of DNA present given the same primer set is being used.
Food Packaging Studies
See materials and methods in McInnis et al. 2018.23
Nuclease Coatings
AMP-7 Antibacterial Testing: A commercial polyurethane (Minwax oil-modified water-based polyurethane) was used for coating preparation and duralar sheets as the substrate. AMP-7 was incorporated at 3% total solids and ground into the coating with milling beads for 30 minutes, and an artist paint brush was used to coat duralar cutouts 1 inch in diameter. E. coli K12 was inoculated into 5 ml TSB media and incubated overnight. Twenty microliters was applied to either the AMP-7 or no coating control duralar cutouts, allowed to dry, and incubated overnight at 37 °C. The cutouts were then placed inoculated side down onto a TSA plate, continued to incubate at 37 °C and monitored every
24 hours for growth.
DNase Plasmid Digest: The following experiments were conducted with dry (lyopholized) DNase (Sigma 10104159001 DNASE I, GRADE II, from bovine pancreas). Unless otherwise indicated pAMP was diluted to 0.01 µg/µL in sterile ddH2O and 5 µL (or 10 µL) was added to each well and cured at 50 °C for 30 min then put at room temperature. Plates were allowed to incubate at room temperature for 2 hours, 15 µL of sterile water was added to each well and the contents visualized using gel electrophoresis and UV light exposure. DNase coating was prepared by adding 150 µL of 10 mg/mL DNase stock to 500 mg of polyurethane (or 300 ul plus 1ml). The DNase coating was drawn down on duralar sheets at 6 mil wet thickness and cured overnight at room temperature. For gel electrophoresis, pAMP loading was 5µL of a 0.02 µg/µL solution (final of 0.1 µg). Image was prepared at 1.5% DNase
E. coli Transformation: E. coli K12 (Presque Isle Culture collection) cells were made competent by calcium chloride treatment. Briefly, E. coli was inoculated from an agar plate into 5 mL of TSB broth, incubated at 37 °C for 4 hours, and centrifuged at 4,000 rpm for 10 min. The supernatant was removed, then the cells were resuspended in 500 µL 50 mM calcium chloride and placed on ice or stored at 4 °C until use. The cutouts were covered with a petri dish lid and incubated at room temperature for 2 hours. Five microliters of pAMP (challenged with 16 nanograms of pAMP plasmid DNA, by diluting pAMP to 3.2 ng/µL and adding 5 µL to each cutout in triplicate) was added to the coated surface and incubated at 30 °C for 2 hours. Ten microliters of cold competent E. coli K12 cells was added to the coatings, which were placed on ice for 15 minutes, followed by 42 °C water bath for 2 min, and back on ice for an additional 2 min. One hundred and ninety microliters of SOC media was added, and 50 µL was plated onto LB/AMP plates.
Waterborne coatings, in particular, have been an important market for biocide manufacturers. However, the necessity to preserve these products from spoilage is challenging as regulations have increased on the type and level of biocide that can be used. This includes the use of formaldehyde releasers, growing restrictions on levels of isothiazolinones, as well as heavy metals such as chromium. Companies using these products have struggled to find ways to meet the new regulations while also offering quality, microbe-free products to their customers. The coatings industry’s focus on traditional biocides has led to predictions that no new actives are on the horizon, but bio-based additives offer solutions to turn that trend around. In cases where bioadditives alone are not sufficient to eliminate all microbial contamination, they can be used synergistically to lower the levels of traditional biocides needed. By applying biotechnology and molecular biology approaches, those levels can be lowered further. In addition, by capitalizing on the potential of these safe and effective bioadditives, manufacturers can expand into markets they might not have yet considered. Herein is presented the application of biotechnology for the preservation of coatings to achieve these goals.
Introduction
Waterborne coatings have been a lucrative market for biocide manufacturers over the last several decades with the move away from solvent-based systems and the increased demand for low-VOC coatings. With that move came an increased need to control microbial contamination and spoilage in water-based systems because these aqueous environments and organic nutrient sources offer an environmental niche well-suited for microbial growth. However, health concerns regarding the biocides used, coupled with increasing restrictions on the types and concentrations of traditional biocides, have increased the demand for development of novel and less toxic preservatives.1-3
Different microbial groups are responsible for negative effects on coating properties. For this reason, the biocides used routinely differ depending on their use in liquid in-can versus dry-film products. Bacteria are of greater concern for in-can preservation, while fungi and algae are the greatest threat to dry coatings, and both microbes and macrofauna are important when considering antifouling systems. In aqueous systems, microbial growth can impact the coating quality by hydrolyzing components, decreasing pH, producing gases, creating foul odors, discoloring the in-film product, and decreasing viscosity.4-5
Traditional biocides approved for in-can preservation typically include formaldehyde releasers, isothiazolinone derivatives, as well as brominated and other halogenated compounds. These are often combined with classes such as carbamates, quaternary amines, phenyl urea derivatives, and heavy metals for dry-film preservation and antifouling properties. Mechanisms of biocidal activity vary, but include effects due to alkylating agents, crosslinkers, electrophilic substances, membrane disruptors, as well as releasers of free radicals and reactive oxygen species.4 Some of these have more direct impact on humans (e.g., alkylating agents and crosslinkers), while others, such as the isothiazolinones, have lower direct toxicity to humans, but may cause sensitization following continued exposure.6-8 These and other health risks, as well as potential environmental impacts, have prompted many countries to restrict the levels of use and/or require special labeling.9-10
Table 1 lists some of the common traditional biocides. Many of the in-can preservative agents contain either 2-methyl-4-isothiazolin-3-one (MIT), or one of its isothiazolinone derivatives, making up brand name products like Kathon 1.5 and Rocima, all of which are subject to growing regulations restricting their usage and levels. From the health concerns with formaldehyde releasers and skin sensitizers, to the United States reaction following the increased standards in Europe [like REACH and the 2013 EU Biocides regulations (BPR) act], and calls by some to avoid biocide agents in building products altogether, these factors continue to put pressure on the industry.2, 11 Novel approaches are needed to meet these mounting regulatory hurdles, while still providing the needed protection in susceptible water-based coatings.
The use of rapid molecular techniques are described herein to select bio-based molecules which are safe and effective at controlling microbial contamination and can be used synergistically with existing biocides, with the goal of reducing the levels of such traditional biocides. Examples of bio-based molecules include enzymes like lysozyme and glucose oxidase, small peptides similar in activity to natural antimicrobial peptides like defensins, but smaller in molecular weight, and even whole cells containing such molecules. By retaining the natural biological function (in either direct antimicrobial activity or synergistic activity with a traditional biocide) it is possible to utilize these molecules in liquid and dry film coating systems.
Experiments and Results
Application of Molecular Techniques to Screen Bio-based Additives as In-Can Preservatives
The small peptides, AMP-6 and AMP-7, are 6-7 amino acids in length (molecular weight under 1000 g/mol) and were selected from a peptide library due to their antimicrobial properties. The library was created using combinatorial synthesis methods to produce over 53 million possible peptides that could be screened for specific antimicrobial activities. Figure 1A shows an example of the iterative amino acid substitution process used to generate the library and minimum inhibitor concentrations (MIC) against Fusarium, a common filamentous fungus.12-16 By altering the peptide sequence, it is possible to “tune” the activity towards specific target microbe groups like bacteria versus fungi and molds. Using the combinatorial chemical approach, progressively lower concentrations are needed to kill the same target microorganism as the peptide chemical structure becomes more defined (which in this case is a 6-amino acid chain). In the case of AMP-6 (a 6-amino acid peptide), greater activity was seen against fungi (as indicated by a lower MIC number) than the bacteria tested (a higher concentration needed). With the addition of just a single amino acid, AMP-7 shows satisfactory activity against both bacteria and fungi (Figure 1B).
Molecular Biology Techniques Drop Test Times to Minutes vs. Weeks
The use of molecular-based microbial viability assays, such as the XTT assay, allows for rapid assessment of the effects of candidate antimicrobials.17 XTT is a tetrazolium dye (2,3-bis-(2-methoxy-4-nitro-5-sulfophenyl)-2H-tetrazolium-5-carboxanilide) that is reduced to colored products by metabolically active cells (darker reddish-brown wells) versus cells that are heat-killed or treated with ethanol, which remain the lighter yellow color (Figure 2A). The difference in the absorbance spectrum can be measured using a spectrophotometer and the viability of the cell population rapidly assessed. Applying these assays, it is possible to screen for an antimicrobial’s selectivity against particular contaminating microorganisms rather than apply broadly toxic biocides. Figure 2B shows that the concentrations of bioadditives needed to control different contaminating strains of Pseudomonas can vary drastically. For P. putida, several agents were effective at various concentrations (as indicated by higher % reduction in metabolism), while P. aeruginosa required the higher test concentration (blue bars) of the tested bio-additive.
Synergistic Activity with Existing Traditional Biocides
Based on the initial XTT screening results, two common biocides were evaluated for synergistic activity with these bio-based agents: an isothiazolinone (MIT), and the formaldehyde releaser, 1,3-dihydroxymethyl-5,5-dimethylhydantoin (DMDM) (Figure 3). A mixed inoculation test panel of six common bacterial spoilage agents, including several Pseudomonads and other gram-negatives like Enterobacter and a gram-positive Bacillus, was used to evaluate the bioadditives and biocides. In the first column, these agents are shown to be effective alone at 15 ppm (15 ppm is the cutoff for labeling as a skin sensitizing agent versus the more stringent EUH208 “May produce an allergic reaction”). At 15 ppm little inhibition was seen in this assay (only around 15% reduction in metabolism), but when combined with particular bio-based additives, an increase in the effectiveness was seen (from 15% up to over 70%, which is comparable to what is seen in heat-killed cells). Particularly with the glucose oxidase synergistic activity an effective kill rate at 15 ppm is achieved. Glucose oxidase catalyzes the oxidation of glucose and subsequent release of hydrogen peroxide that can induce cellular damage.18, 20 Glucose oxidase is listed by the Food and Drug Administration as generally recognized as safe (GRAS) for various intended uses and is frequently used in the food industry.19 It is produced on commercial scale and available in bulk quantities. This apparent synergistic activity was confirmed by testing varying concentrations of glucose oxidase and MIT in the XTT assay. The responses of the test panel bacteria to various combinations of glucose oxidase and MIT are displayed in the heatmap (Figure 4A), which shows the relative response as a color from red (highest percent growth inhibition) to green (lowest percent growth inhibition). To test whether antagonism or synergy exists between these two compounds, the responses from glucose oxidase and MIT combinations were used to determine the synergy score using the zero interaction potency (ZIP) method, which returns a score based on the deviation of the actual response from the expected response.21 These scores are visualized for each combination in a contour plot (Figure 4B), which suggests that the two antimicrobials generally interact in a synergistic manner. Regions of synergism exist for concentrations of glucose oxidase between 50 ppm and 500 ppm and for concentrations of MIT ranging from below 0.15 ppm to 1 ppm, as well as the greatest synergy seen for concentrations of glucose oxidase between 10 ppm and 1,000 ppm and for concentrations of MIT ranging from below 7 ppm to 15 ppm. This suggests that the glucose oxidase and MIT are complementing each other’s function at both low and high concentrations allowing for lower levels of MIT to show effective antimicrobial activity.
Evaluation of Glucose Oxidase Synergy with Traditional Biocides Using ASTM D2574
The antimicrobial activity of glucose oxidase, detected using the rapid molecular techniques, was further tested as an in-can coatings preservative using the traditional testing method, ASTM D2574 Paint Spoilage standard.22 By using the above molecular methods to rapidly screen target microbes, it is possible to select the most promising candidates to classic tests like ASTM D2574.
For this testing, a biocide-free acrylic latex was used (see Appendix - Materials and Methods for formulation). The same spoilage agent test panel used for the XTT assay was used to challenge the acrylic latex; the samples were monitored over seven days. The selected dosage of glucose oxidase for the ASTM D2574 procedure was based on the results from the XTT assay, where 50 ppm showed synergy with both MIT and DMDM while 5 ppm was only synergistic with DMDM. Figure 5 shows example plates from bio-based additive versus traditional biocides on the final day. Over the 7-day monitoring period the biocide-free latex controls scored a 4 for each day (complete overgrowth).
While the recommended dosages of MIT and DMDM achieved a 0 (no growth) on day 1, the lower doses of 15 ppm were less effective with DMDM still showing a 4 on day 7 and MIT not achieving a 0 until day 5. However, in the ASTM method using the biocide-free latex, the lowest test concentration of glucose oxidase (50 ppm) achieved a 0 on day 1 and when combined with 15 ppm MIT or DMDM a 0 was also achieved on day 1 (Table 2).
While it is a positive result that glucose oxidase alone showed such high activity, further studies are being conducted at lower concentrations of glucose oxidase in order to characterize any synergistic activity with traditional biocides in the acrylic latex.
Molecular Approaches for Detecting and Eliminating Paint Spoilage Agents
While classic testing methods like ASTM D2574 identify broadly toxic agents that kill all challenge organisms, they fail to selectively identify agents effective against particular strains. By using DNA sequencing methods, it is possible to identify specifically which bacteria are killed by particular bio-based additives for precision treatments (Figure 6A).
Real-time quantitative polymerase chain reaction (PCR) was used for direct analysis of paint for microbial contamination and identification of the specific contaminant. The graph in Figure 6B shows the use of selective probes to identify particular bacterial species. The fluorescence increases as the strain-specific probes successfully amplify the target DNA. These methods can also be used to determine the levels of contamination as well. Figure 6C shows a dose response curve using Enterobacter, which indicates the changes in cycle number needed to see amplification at the different concentrations. By identifying the particular level of bacterial contamination and treating accordingly, the need to use high levels of broadly toxic biocides in products decreases.
Expanded Markets Using Bio-based Additives: Food Safety and Combating Antibiotic Resistance
With the low toxicity bio-based additives, some of which have GRAS status for various applications, manufacturers can expand into new markets beyond in-can coating preservation - one example being food packaging. A rapid screening technique based on inhibition of microbial colony formation on agar plates was used to find active agents against E. coli and explore any potential synergy between active agents.23 Figure 7A shows an image of small 2.5 cm polyethylene terephthalate (PET) cutouts coated with polyvinyl alcohol (PVA) incorporating bioadditives used to test for contaminant inhibition (in this case E. coli) on agar plates. With increasing concentrations of each agent alone, inhibition of growth was observed, as measured by the reduction in the number of visible colonies under the plastic. By combining the two agents, a further reduction in the concentration of each agent needed was achieved. Following these results, a lab-scale test was developed using food simulant patties (microbial growth media with cell growth color indicator) that can be challenged with the test microbe and then vacuum sealed with either bio-additive coated sealing plastic itself or sandwiched between inserts with bio-additive coatings to prevent foodborne contamination (Figure 7C). As seen in Figure 7B, at an optimal dose of bio-additive in the packaging coating, the viability of the E. coli is reduced to zero (no colonies).
Other examples of expanded markets include products to help combat a growing global healthcare concern: antibiotic resistance. Not only is there a need for safe and effective antimicrobials (such as the bio-based agents described above), but also methods to combat transfer of antibiotic resistance (as a cell is lysed, the DNA spills out and can be taken up by other cells).
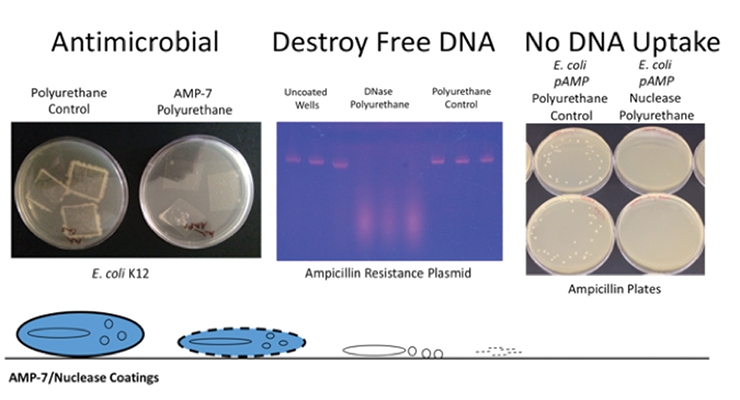
Enzymatic coating surfaces have been successfully created that can breakdown and destroy cell- free DNA spilled from a cell before it can be taken up by other cells.24 The top left panel in Figure 8 shows the effective killing of E. coli by AMP-7 (3% total solids), as indicated by lack of bacterial colony growth around the edges of the AMP-7 polyurethane cutout versus the control when placed on contaminated agar plates. The middle panel shows the destruction of DNA by a polyurethane coating containing the nuclease, DNase I (1.5% total solids). The DNA used here carries a gene for resistance to ampicillin, a beta-lactam antibiotic related to penicillin. The DNA was applied to the surface and allowed to dry. Following exposure, the DNA was resuspended from the coating surface and analyzed. On both ends of the image in Figure 8 are three wells each of coating controls (either the substrate itself or the polyurethane coating). In the middle are wells with the DNA exposed to the nuclease-containing coating. Distinct bands are no longer detected when exposed to the nuclease-containing coating surface, though the smear of the smaller breakdown products can be seen, indicating that the DNA has been destroyed. Furthermore, on the right in Figure 8, it was demonstrated that following exposure of the ampicillin resistance gene containing DNA to the nuclease-coated surface, no transfer of antibiotic resistance to E. coli occurs. The agar plates in this image have ampicillin added to them, and only E. coli that were able to take the DNA directly from the surface are able to grow (n=3: 36, 37, and 26 colonies). However, E. coli from the nuclease-coated surface did not develop resistance to ampicillin because the DNA was destroyed, resulting in no colony growth (n=3: 0, 0, and 0 colonies). The bottom of Figure 8 sums up the overall process. After the bacteria are lysed and killed by AMP-7, the free DNA is degraded by nucleases in the coating, and the resistance gene is not passed.
Further Coating Manufacturing Applications to Reduce Biocide Levels
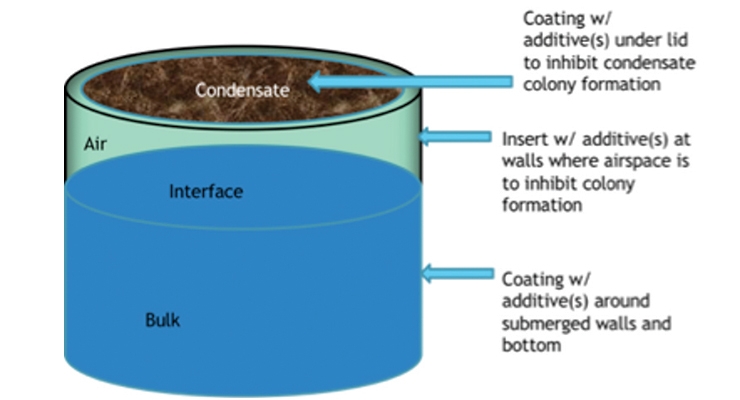
In the coatings manufacturing arena these technologies can be applied to expand applications and market areas even further, such as surgical strikes by inclusion of selected bioadditives along the coating manufacturing process line to decrease the need to incorporate high levels of biocides in the final product (Figure 9A). By using the previously mentioned molecular methods, the source contaminant can be rapidly identified and equipment lines and materials treated accordingly with bio-additives at the point of contamination. By utilizing biomolecules, like enzymes, and the tunability of AMPs, the targeted use of these agents within the plant becomes possible. That protection can potentially continue from the plant to the consumer without an added in-can biocide by coating the can itself with bio-additive protectants as shown in Figure 9B. Greater access to oxygen at the walls and the paint/headspace interface makes these active sites of microbial growth, and condensation buildup on the lid makes this another potential problem site to target.
Conclusion
In conclusion, given the growing regulatory challenges facing traditional biocide usage, alternatives are needed to 1) address the increased regulations and 2) expand upon existing markets. One such approach to address those issues is the use of rapid molecular methods to identify bio-based additives such as peptides and enzymes for selective contaminant targeting. By using these methods, agents were identified that have either inherent antimicrobial activity, like AMP-6 and AMP-7, or GRAS enzymes like glucose oxidase. In instances where the bio-based biocide is not broadly antimicrobial on its own, it is shown here that such bio-based molecules facilitate the activity of traditional biocides like MIT and allow for lower levels of biocide use.
Acknowledgements
Aayushma Kunwar, Kyle Wilhelm and Earnest Couch, student interns from William Carey University, Hattiesburg, MS, are thanked for their assistance with these projects.
References
1. Cresswell, T.; Richards, J. P.; Glegg, G. A.; Readman, J. W., The impact of legislation on the usage and environmental concentrations of Irgarol 1051 in UK coastal waters. Marine Pollution Bulletin 2006, 52 (10), 1169-1175.
2. Brinch, A.; Hansen, S.; Hartmann, N.; Baun, A., EU regulation of nanobiocides: challenges in implementing the biocidal product regulation (BPR). Nanomaterials 2016, 6 (2), 33.
3. White Paper: Antimicrobial Building Products Should Be Avoided Whenever Possible. http://perkinswill.com/news/perkinswill-white-paper-antimicrobial-building-products-should-be-avoided-whenever-possible (accessed December 2017).
4. Sauer, F., Microbicides in Coatings. Vincentz Network: Hanover, Germany, 2017.
5. Winkowski, K., Efficacy of in can preservatives. European coatings journal 2001, (1-2), 87-91.
6. Chan, P. K.; Baldwin, R. C.; Parsons, R. D.; Moss, J. N.; Stiratelli, R.; Smith, J. M.; Hayes, A. W., Kathon biocide: manifestation of delayed contact dermatitis in guinea pigs is dependent on the concentration for induction and challenge. Journal of investigative dermatology 1983, 81 (5), 409-411.
7. Schwensen, J. F.; Menné, T.; Andersen, K. E.; Sommerlund, M.; Johansen, J. D., Occupations at risk of developing contact allergy to isothiazolinones in D anish contact dermatitis patients: results from a D anish multicentre study (2009–2012). Contact Dermatitis 2014, 71 (5), 295-302.
8. Schwensen, J. F.; Lundov, M. D.; Bossi, R.; Banerjee, P.; Gimenez-Arnau, E.; Lepoittevin, J. P.; Lidén, C.; Uter, W.; Yazar, K.; White, I. R., Methylisothiazolinone and benzisothiazolinone are widely used in paint: a multicentre study of paints from five E uropean countries. Contact Dermatitis 2015, 72 (3), 127-138.
9. Bollmann, U. E.; Fernández-Calviño, D.; Brandt, K. K.; Storgaard, M. S.; Sanderson, H.; Bester, K., Biocide runoff from building facades: degradation kinetics in soil. Environmental science & technology 2017, 51 (7), 3694-3702.
10. Lundov, M. D.; Kolarik, B.; Bossi, R.; Gunnarsen, L.; Johansen, J. D., Emission of isothiazolinones from water-based paints. Environmental science & technology 2014, 48 (12), 6989-6994.
11. REACH Legislation. https://echa.europa.eu/regulations/reach/legislation (accessed 08/09/19).
12. McDaniel, C.S., Anti-fouling Coatings and Coatings,
U.S. patent application 14/097,128, July 9, 2015.
13. McDaniel, C.S., Antifungal Coatings and Coatings, U.S. patent 8,497,248, July 30, 2013.
14. McDaniel, C.S., Coating Compositions having Peptidic Antimicrobial Additives and Additives of Other Configurations, U.S. patent 8,618,066, December 31, 2013.
15. McDaniel, C.S., Antifungal Coatings and Coatings, U.S. patent 7,932,230, April 26, 2011.
16. McDaniel, C.S., Anti-fouling Coatings and Coatings, U.S. patent 7,939,500, May 10, 2011.
17. XTT Cell Proliferation Assay Kit Instruction Manual. American Type Culture Collection: Manassas, VA, 2011.
18. Hodges, T. W.; Kemp, L. K.; Mcinnis, B. M.; Hurt, J. D.; McDaniel, S.; Rawlins, J. W.; Wilhelm, K. L., Proteins and Peptides: as Replacements for Traditional Organic Preservatives: Part I. CoatingsTech 2018, 15 (4).
19. Bankar, S. B.; Bule, M. V.; Singhal, R. S.; Ananthanarayan, L., Glucose oxidase—an overview. Biotechnology advances 2009, 27 (4), 489-501.
20. Tiina, M.; Sandholm, M., Antibacterial effect of the glucose oxidase-glucose system on food-poisoning organisms. International Journal of Food Microbiology 1989, 8 (2), 165-174.
21. He, L.; Kulesskiy, E.; Saarela, J.; Turunen, L.; Wennerberg, K.; Aittokallio, T.; Tang, J., Methods for High-throughput Drug Combination Screening and Synergy Scoring. Methods Mol Biol 2018, 1711, 351-398.
22. ASTM Standard D2574-16, Standard Test Method for Resistance of Emulsion Coatings in the Container to Attack by Microorganisms. ASTM International: West Conshohocken, PA, 2016.
23. McInnis, B. M.; Hodges, T. W.; Kemp, L. K.; Hurt, J. D.; McDaniel, S.; Kunwar, A., Bio-based Antimicrobial Food Packaging Coatings. CoatingsTech 2018, 15 (9), 36-43.
24. Ellison, C. K.; Dalia, T. N.; Vidal Ceballos, A.; Wang, J. C.-Y.; Biais, N.; Brun, Y. V.; Dalia, A. B., Retraction of DNA-bound type IV competence pili initiates DNA uptake during natural transformation in Vibrio cholerae. Nature Microbiology 2018, 3 (7), 773-780.
25. Reed, J. D.; Edwards, D. L.; Gonzalez, C. F., Synthetic Peptide Combinatorial Libraries: A Method for the Identification of Bioactive Peptides Against Phytopathogenic Fungi. Molecular Plant-Microbe Interactions 1997, 10 (5), 537-549.
26. ZymoBIOMICS DNA/RNA Miniprep Kit. www.zymoresearch.com: Ver. 1.2.0.
27. Lee, C. S.; Wetzel, K.; Buckley, T.; Wozniak, D.; Lee, J., Rapid and sensitive detection of Pseudomonas aeruginosa in chlorinated water and aerosols targeting gyrB gene using real-time PCR. Journal of applied microbiology 2011, 111 (4), 893-903.
28. Preston, K. E.; Radomski, C. C.; Venezia, R. A., Nucleotide Sequence of the ChromosomalampC Gene of Enterobacter aerogenes. Antimicrobial agents and chemotherapy 2000, 44 (11), 3158-3162.
Appendix Materials and Methods
Antimicrobial Peptide Library
See materials and methods in Reed et al. 200725 and patent numbers patent numbers 5,602,097, 5,885,782, and 6,020,312.
XTT Assay
See materials and methods in Hodges et al. 2018.18
XTT Assay for Synergy with MIT and DMDM
See materials and methods in Hodges et al. 2018.18 For experiments involving the combination glucose oxidase and MIT, the interaction between the two additives (i.e., synergistic, additive, antagonistic) was evaluated using the zero interaction potency (ZIP) model with the R package synergyfinder.21,23
ASTM D2547 Using Acrylic Latex
Antimicrobial activity of traditional and bio-based biocides were assessed using the ASTM method D254722 using a custom acrylic latex (50 wt% solids, pH = 8, Tg = 18 °C) without biocide. Twenty-five milliliters of liquid acrylic latex was dispensed into sterile glass jars, and appropriate quantities of glucose oxidase, MIT, or DMDM were hand-stirred into the coating using sterile glass rods. Sterile swabs were dipped into each sample and used to streak Tryptic Soy Agar (TSA) plates to ensure that the samples were free of bacterial contamination. Bacterial cultures of Pseudomonas aeruginosa, Pseudomonas putida, Pseudomonas fluorescens, Bacillus cereus, Alcaligenes faecalis, and Enterbacter aerogenes were grown in 5 mL Tryptic Soy Broth (TSB) at 30 °C with agitation for 24 hours. Sterile loops full of each culture were passed into new 5 mL of TSB, which were incubated at 30 °C with agitation for 24 hours. Equal volumes of the cultures were passed once more, with a final incubation at 30 °C with agitation for 24 hours. The cultures were mixed, and 250 µL of the bacterial mixture was dispensed into each latex sample. Sterile swabs were dipped into each sample and used to streak duplicate TSA plates 1, 3, 5, and 7 days following inoculation. The TSA plates were incubated at 30 °C for 7 days, after which each plate was scored using the score system mandated by the ASTM
D2547 procedure:
0 - no bacterial recovery
1 - trace contamination (1 to 9 colonies)
2 - light contamination (10 to 99 colonies)
3 - moderate contamination (> 100 distinct colonies)
4 - heavy contamination (continuous smear of growth,
colonies have grown together and are indistinguishable)
Isolate Identification using Molecular Analysis
Following inoculation of biocide-free acrylic latex paint or bio-based additive treated paint according to ASTM D2547, bacterial colonies were steaked for isolation on TSB agar. Isolated colonies were transferred into microcentrifuge tubes containing 100 µL of chelex solution (BioRad Instagene Matrix). The tubes were vortexed for 15 seconds and placed in a 70 °C water bath for 1 hour, with vortexing repeated every 15 minutes. Samples were then centrifuged at 12,000xg for 2 min and either used directly in polymerase chain reaction (PCR) or frozen. PCR primers targeting a portion of the 16S ribosomal RNA (rRNA) gene were used, specifically 8F (5’-AGAGTTTGATCMTGGCTCAG-3’) and 1392R (5’-ACGGGCGGTGTGTACA-3’). GE Healthcare Illustra PureTaq PCR beads were used for PCR amplification with the following conditions: Per reaction vial: 2 µL 8F primer, 2 µL 1392R primer, 2 µL template DNA, and 19 µL sterile water. Thermal cycler conditions were 94 °C for 5 min followed by 30 cycles of 94 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min followed by 72 °C for 7 min. Following completion of PCR, Invitrogen ChargeSwitch-Pro PCR Cleanup Kit was used to purify the amplification products. PCR products containing the amplified region of the 16S rRNA gene were then sent for sequencing at Eurofins Genomics (Louisville, KY). Upon receipt of the sequence data, sequences were searched against the NCBI Genbank using BLAST (Basic Local Alignment Search Tool). This allows for strain identification based on sequence similarity to deposited 16S rRNA gene sequences.
Molecular Paint Analysis Using Quantitative Real-time PCR
Direct DNA extraction from paint: Five types of commercially available DNA extraction kits were tested for effectiveness in isolating amplifiable bacterial DNA from latex acrylic paint (AxyPrep Mag Tissue-Blood gDNA, Invitrogen PureLink, Microbiome DNA Purification Kit, Invitrogen PureLink Genomic DNA Mini Kit, and ZymoBiomics DNA Mini Kit). Off-the-shelf Sherwin Williams Harmony EgShel was inoculated with overnight cultures of set bacteria, in this example Enterobacter aerogenes, followed by DNA extraction using the commercially available kits.26 Amplification using universal bacterial 16S primers (16F and 16R) and standard end-point PCR showed amplification as indicated by appropriate size bands in gel electrophoresis.
Reaction Conditions and PCR Primers: Quantitative Real-time PCR conditions were optimized using primers targeting functional genes in three species from the ASTM challenge method (Pseudomonas aeruginosa, Enterobacter aerogenes, and Bacillus cereus). Where necessary, primers were designed using the National Center for Biotechnology Information Primer-BLAST function. An Eppendorf RealPlex thermal cycler and SYBR Green were used for detection of amplified DNA (Applied Biosystems™ PowerUp™ SYBR™ Green Master Mix). Primers were synthesized by Eurofin Operon (Louisville, KY).
Primers sequences used:
Pseudomonas aeruginosa: gyrB gene (DNA gyrase, subunit B)27
PA-722 primer set: 722F:5’-GGCGTGGGTGTGGAAGTC-3’
and 722R: 5’-TGGTGGCGATCTTGAACTTCTT-3’)
Enterobacter aerogenes: Designed based on beta-lactamase
AmpC gene28
EA-PP5 primer set: PP5F:5’-GACTACGCCTGGGGTTATCG-3’
PP5R: 5’-CGCTGTTCATATTCGCCAGC-3’
Typical reaction mixtures: Primers were diluted to 20 µM stocks in sterile ddH20 and 0.5 µL added per 20 µL reaction for a final concentration of 500 nM. DNA concentrations varied between experiments: extracted genomic DNA was used at 2 µL in initial experiments and, if quantitated, it was added at 0.5 µL per 20 µL reaction (pg or ng range). Example of a typical 20 µL reaction setup is as follows:
Per 20 µL reaction
0.5 µL Forward Primer
0.5 µL Revers Primer
0.5 µL DNA (up to 2 µL)
7.5 µL sterile H2O
10 µL SYBR Green Master Mix
Thermal Cycler Conditions: 50 °C for 2 min, 95 °C for 2 min followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min with fluorescence measured at 520 nM at the end of each cycle. Cycle threshold values (ct values) were calculated automatically by the instrument. This is the number of cycles taken for the fluorescence signal to cross background noise level and is inversely proportional to the amount of DNA present given the same primer set is being used.
Food Packaging Studies
See materials and methods in McInnis et al. 2018.23
Nuclease Coatings
AMP-7 Antibacterial Testing: A commercial polyurethane (Minwax oil-modified water-based polyurethane) was used for coating preparation and duralar sheets as the substrate. AMP-7 was incorporated at 3% total solids and ground into the coating with milling beads for 30 minutes, and an artist paint brush was used to coat duralar cutouts 1 inch in diameter. E. coli K12 was inoculated into 5 ml TSB media and incubated overnight. Twenty microliters was applied to either the AMP-7 or no coating control duralar cutouts, allowed to dry, and incubated overnight at 37 °C. The cutouts were then placed inoculated side down onto a TSA plate, continued to incubate at 37 °C and monitored every
24 hours for growth.
DNase Plasmid Digest: The following experiments were conducted with dry (lyopholized) DNase (Sigma 10104159001 DNASE I, GRADE II, from bovine pancreas). Unless otherwise indicated pAMP was diluted to 0.01 µg/µL in sterile ddH2O and 5 µL (or 10 µL) was added to each well and cured at 50 °C for 30 min then put at room temperature. Plates were allowed to incubate at room temperature for 2 hours, 15 µL of sterile water was added to each well and the contents visualized using gel electrophoresis and UV light exposure. DNase coating was prepared by adding 150 µL of 10 mg/mL DNase stock to 500 mg of polyurethane (or 300 ul plus 1ml). The DNase coating was drawn down on duralar sheets at 6 mil wet thickness and cured overnight at room temperature. For gel electrophoresis, pAMP loading was 5µL of a 0.02 µg/µL solution (final of 0.1 µg). Image was prepared at 1.5% DNase
E. coli Transformation: E. coli K12 (Presque Isle Culture collection) cells were made competent by calcium chloride treatment. Briefly, E. coli was inoculated from an agar plate into 5 mL of TSB broth, incubated at 37 °C for 4 hours, and centrifuged at 4,000 rpm for 10 min. The supernatant was removed, then the cells were resuspended in 500 µL 50 mM calcium chloride and placed on ice or stored at 4 °C until use. The cutouts were covered with a petri dish lid and incubated at room temperature for 2 hours. Five microliters of pAMP (challenged with 16 nanograms of pAMP plasmid DNA, by diluting pAMP to 3.2 ng/µL and adding 5 µL to each cutout in triplicate) was added to the coated surface and incubated at 30 °C for 2 hours. Ten microliters of cold competent E. coli K12 cells was added to the coatings, which were placed on ice for 15 minutes, followed by 42 °C water bath for 2 min, and back on ice for an additional 2 min. One hundred and ninety microliters of SOC media was added, and 50 µL was plated onto LB/AMP plates.